If you’ve ever watched a prime time crime drama such as CSI, you probably recall a scene in which a forensic analyst used a computer to troll through thousands of snippets of DNA, looking for a match between a crime scene and a suspect. Real life doesn’t happen quite like it does on television, but the gist is the same. Genetics is inherently a comparative science. Whether you’re trying to identify a suspect, a genetic disease or a long-lost relative, it involves comparing one genome to another, hunting for telling similarities or variabilities among billions of letters of DNA.
Image: NIH
But while finding a relative or a crime suspect usually involves looking at just a few snippets of a person’s genetic code, problems such as identifying the genetic variant responsible for a disease require churning through a lot more data. Even with all the fancy maths designed to help scientists do this, making sense of all that data is still a big challenge. It’s also exactly the kind of problem that artificial intelligence is designed to solve.
This week, Google released a tool called DeepVariant that uses deep learning to piece together a person’s genome and more accurately identify mutations in a DNA sequence.
Built on the back of the same technology that allows Google to identify whether a photo is of a cat or dog, DeepVariant solves an important problem in the world of DNA analysis. Modern DNA sequencers perform what’s known as high-throughput sequencing, returning not one long read out of a full DNA sequence but short snippets that overlap. Those snippets are then compared against another genome to help piece it together and identify variations. But the technology is error-prone, and it can be difficult for scientists to distinguish between those errors and small mutations. And small mutation matter. They could provide significant insight into, say, the root cause of a disease. Distinguishing which base pairs are the result of error and which are for real is called “variant calling”.
There are already tools out there to help scientists do this. The most widely used is GATK, a human-engineered algorithm that applies statistics to suss out where sequencing machines most often make mistakes. DeepVariant, though, leverages neural network technology to build something more accurate than anything else in existence. Last year, it won first place in an FDA contest aimed at improving the accuracy of genetic sequencing.
Neural networks are so named because they’re somewhat analogous to how neurons work in the brain. Each layer of the network deals with a progressively more complex problem. To use an image-recognition AI to build an accurate DNA sequence, Google’s team turned DNA-sequencing data into an image. The As, Ts, Cs and Gs that make up a genetic code, for example, became visually represented as red. Researchers then trained their network on millions of sequenced genomes and high-throughput reads, teaching it which things to weigh more heavily and which to ignore.
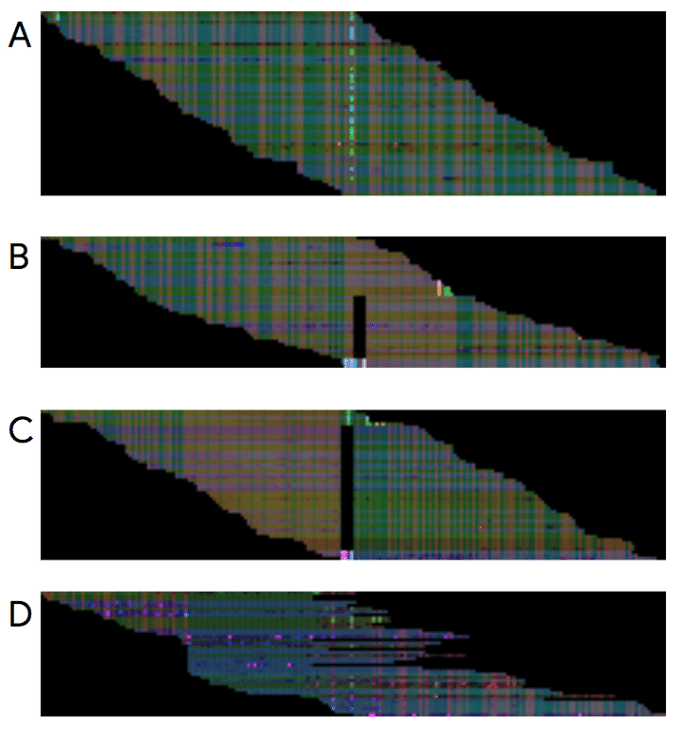
Image: Google
The resulting algorithm can sort actual mutations from errors with more accuracy than any system before. Initially, the images were made up of just three colours, or three layers of data. But the latest version released this week contains seven, making it even more finely tuned. It was released as open-source software, which outside researchers can use and even augment.
DeepVariant is by no means 100 per cent accurate. But its success is representative of the impact machine learning is poised to have on genomics. The scale and complexity of genomic data is immense. Machines might be just what we need to make sense of it.
