The history of AI computing in recent decades follows a somewhat familiar script: brief bursts of industry-shifting breakthroughs followed by months or years of smaller incremental change with a fair share of controversy peppered in between. Today’s one of those watershed moments.
Alphabet-owned Deepmind on Thursday announced it’s releasing a database of predictions for virtually every protein currently known to science, an advancement that’s expected to significantly fast-track drug development and critical advancements in new technologies. The expanded database revealed this week increases the number of known, cataloged proteins included in Deepmind’s database by over 200x, from 1 million structures to around 200 million structures.
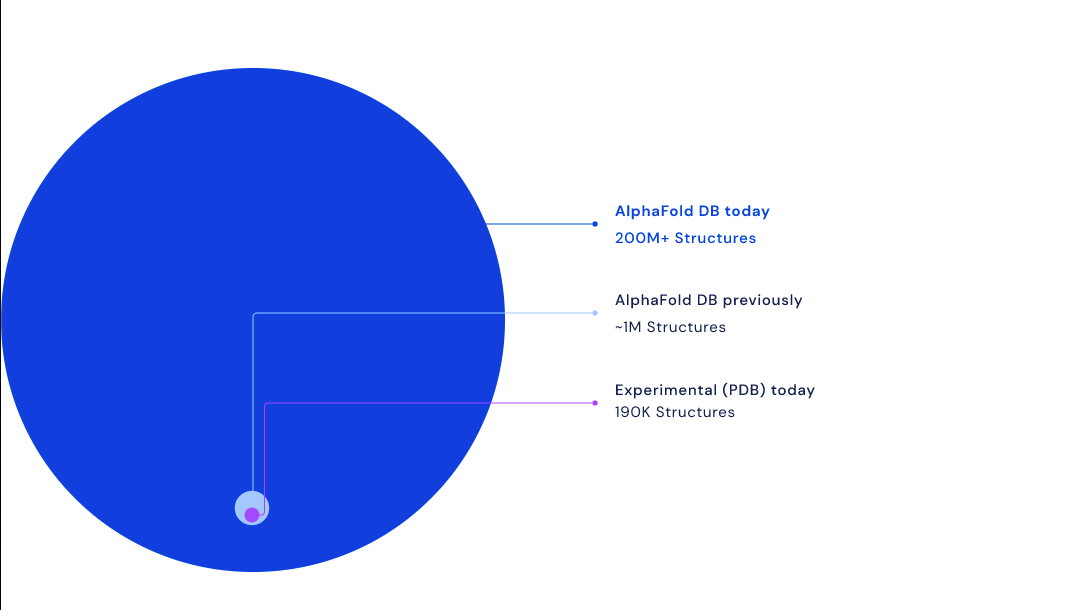
Those predictions come via Deepmind’s AlphaFold AI software. Back in 2020, AlphaFold proved it could predict the shape of certain protein structures and create 3D models with unprecedented accuracy. Deepmind began publishing some of these structures on this open database last year, starting with the known structures of 20 species and 98% of all human proteins. Deepmind believes this week’s hefty expansion, which includes predicted structures for plants, bacteria, animals, and other organisms, could create new opportunities for scientists to advance research needed to address sustainability issues and food scarcity. Deepmind’s making all of the structures available for bulk download through Google’s Cloud Public Datasets.
Prior to AlphaFold, protein prediction reportedly involved time-consuming experimentation involving X-rays, microscopes, and other tools. In a statement, Scripps Research Translational Institute Founder and Director Eric Topol said AlphaFold has reduced the time to accurately predict the structure of a protein from months or years down to mere seconds.
“AlphaFold has already accelerated and enabled massive discoveries, including cracking the structure of the nuclear pore complex,” Topol said. “And with this new addition of structures illuminating nearly the entire protein universe, we can expect more biological mysteries to be solved each day.”
The circles in the image below illustrate the scale of this week’s new additions. While the predicted protein structure for all of the organisms listed increased dramatically since last year, the largest chunk of data involves animals. That’s followed by plants and then shortly after by bacteria.
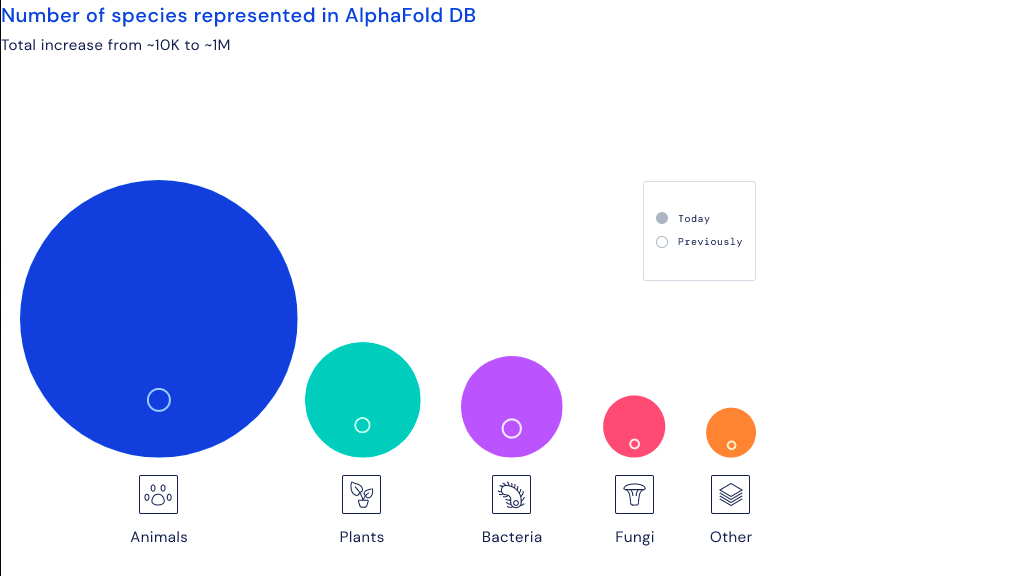
“This comes down to medicine, agriculture, biotech, everything,” European Bioinformatics Institute Director Emeritus Dame Janet Thornton said in a statement. “There are many applications. It’s [the database] like a shop you can go in and just get your favourite protein and look at it, instantly.”
Scientists worldwide have already begun using AlphaFold’s models to advance research in their fields. Naturally, Alphabet’s tried to get in on the action as well. Late last year, the conglomerate announced it had spun off a new company called Isomorphic Labs with the expressed purpose of taking revelations pulled from AlphaFold and using them to discover new pharmaceutical drugs. Ambitiously, Deepmind CEO Demis Hassabis claimed the project could, “reimagine the entire drug discovery process from first principles with an AI-first approach.”